Microbes living on the skin are coming under increasing scrutiny as the cause of the most common type of eczema, called atopic dermatitis. A new pair of studies focus specifically on the bacterium Staphylococcus aureus as playing a role in driving the disorder. The studies – one from the NIH’s National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) and the other from the University of California, San Diego, with NIAMS support – appeared in Science Translational Medicine.
These research papers suggest that a person’s unique skin-dwelling microbiota, the vast array of bacteria, viruses and fungi, may influence susceptibility to atopic dermatitis. Targeting S. aureus could prove to be an effective therapeutic strategy.
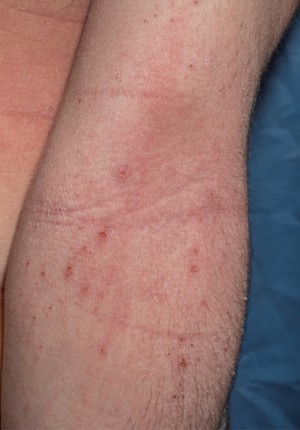
Atopic dermatitis is a chronic, inflammatory condition characterized by patches of red, itchy, inflamed skin. The symptoms ebb and flow, with episodes of high disease activity, or “flares,” alternating with periods of relief. Atopic dermatitis affects as many as 20 percent of children in the United States and often it can start before the age of one. Many children outgrow it, but some continue to have symptoms as adults. There is no cure for the condition, but it can be controlled with lifestyle changes and medicines aimed at reducing inflammation. Gaining a fuller understanding of the factors that underlie atopic dermatitis could lead to better ways to treat it.
With this goal in mind, Heidi H. Kong, M.D., M.H.Sc., head of the cutaneous microbiome and inflammation section at NIAMS, focused her efforts on understanding the role of S. aureus in the disorder. Earlier studies showed that S. aureus is linked to atopic dermatitis, but Dr. Kong wondered if the bacterial population’s genetic diversity changed during a flare.
To find out, her team sampled atopic dermatitis pediatric patients’ skin before, during and after a flare and analyzed the bacteria using advanced gene sequencing techniques. As expected, they saw more S. aureus during flares compared to samples taken before and after and from healthy controls. Researchers also noted a drop in genetic diversity among the bacteria taken from patients having a flare compared to those between flares or from control individuals. Patients experiencing the worst flares carried the least diverse array of bacteria, suggesting a link between disease activity and bacterial diversity.
To see if flare-related bacteria could lead to skin changes, the investigators next administered bacteria from patients experiencing flares to the skin of lab mice. Skin on the mice became inflamed and thick, with the most dramatic response in those receiving samples from patients with the worst symptoms. However, mice treated with bacterial skin samples from healthy control patients did not develop these skin changes, indicating that there is something distinct about S. aureus in atopic dermatitis patients.
“Children with severe eczema have an increased likelihood of developing asthma and hay fever later on,” said Dr. Kong. “Our findings suggest that by understanding the differences in and functions of S. aureus in these children, we may eventually be able to find ways to alleviate the skin-related symptoms, as well as halt the progression of this sequence of allergic disorders.”
In the University of California, San Diego study, Richard Gallo, M.D., and his team, revealed that while bacteria may be part of the problem in atopic dermatitis, they may also be part of the solution. A screening test to identify S. aureus-killing bacteria uncovered two related skin dwelling strains: Staphylococcus epidermis and Staphylococcus hominis. Further research showed that their anti-S. aureus activity comes in the form of several small, previously unidentified protein fragments called antimicrobial peptides. Antimicrobial peptides are key defense molecules found in all forms of life, and their direct action against microbes makes them attractive candidates for development into novel therapeutic agents.
Dr. Gallo’s team created a topical lotion with S. epidermis and S. hominis strains to test their effects on atopic dermatitis patients. Researchers applied the lotion to the inner arm of several volunteers with the condition and saw S. aureus virtually disappear within 24 hours. The same lotion lacking the bacteria had no effect.
“Use of this type of bacterial therapy to fight atopic dermatitis has an advantage over other pharmaceutical approaches in that it specifically targets S. aureus, leaving other types of bacteria unharmed,” said Dr. Gallo. “In addition, antibiotic resistance is less likely to occur because the pathogen is being attacked in multiple ways at the same time.”
The investigators also showed that while the helpful S. epidermis and S. hominis strains are abundant on the skin of healthy people, they are rare on skin of atopic dermatitis patients. This suggests that they provide a first line of defense against the pathogen. Further research is needed to understand why some people’s bodies fail to promote the growth of these helpful types of bacteria.
Dr. Kong’s study was supported by the NIH’s NIAMS (UH3-AR057504, AR-041209, 1ZIA-BC010938-09), National Human Genome Research Institute, National Cancer Institute and National Institute of Allergy and Infectious Diseases (NIAID).
Dr. Gallo’s study was supported by the NIH’s NIAMS (R21-AR067547 and R01-AR06781), NIAID (through the Atopic Dermatitis Research Network, U19-AI117673; R01-AI052453; R01-AI118816) and the National Center for Advancing Translational Sciences (TL1-TR001443). The Rho Federal Systems Division also contributed support.
###
Staphylococcus aureus and Staphylococcus epidermidis strain diversity underlying pediatric atopic dermatitis. Byrd AL, Deming C, Cassidy SKB, Harrison OJ, Ng WI, Conlan S; NISC Comparative Sequencing Program, Belkaid Y, Segre JA, Kong HH. Sci Transl Med. 2017 Jul 5;9(397). pii: eaal4651. doi: 10.1126/scitranslmed.aal4651. PMID: 28679656.
Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Nakatsuji T, Chen TH, Narala S, Chun KA, Two AM, Yun T, Shafiq F, Kotol PF, Bouslimani A, Melnik AV, Latif H, Kim JN, Lockhart A, Artis K, David G, Taylor P, Streib J, Dorrestein PC, Grier A, Gill SR, Zengler K, Hata TR, Leung DY, Gallo RL. Sci Transl Med. 2017 Feb 22;9(378). pii: eaah4680. doi: 10.11.1126 /scitranslmed.aah4680. PMID: 28228596.
